To identify the causes of heterotopia periventricular nodular (HPN) copy number variant (CNV) in patients for whom FLNA sequencing is negative.
Methods:
Screening of 35 patients from 33 pedigrees in an Affymetrix 6.0 microarray led to the identification of one individual with a CNV that disrupted FLNA. FLNA-disrupting CNVs were also isolated from 2 other individuals by multiplex ligation probe amplification. These 3 cases were further characterized by high-resolution oligo array comparative genomic hybridization (CGH), and the precise junctional breakpoints of the rearrangements were identified by PCR amplification and sequencing.
Results:
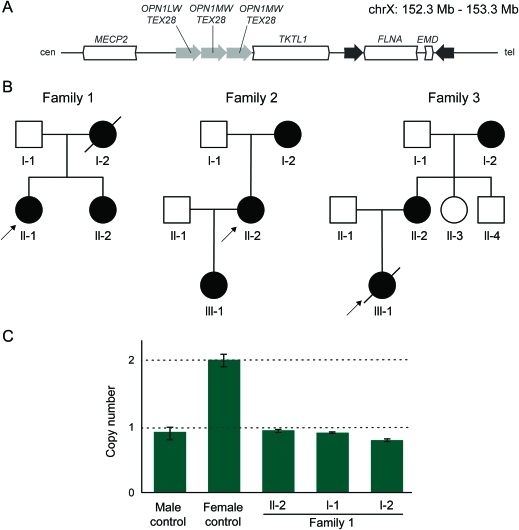
We report 3 cases of PNH caused by non-recurrent genomic rearrangements that disrupt one copy of FLNA. The first individual carried a 113 kb deletion that removes all but the first FLNA exon. A second patient harbored a complex rearrangement that included a deletion of the 3' end of FLNA accompanied by a partial duplication event. A third patient had a 39 kb deletion encompassing the entire FLNA and the neighboring EMD gene. High-resolution trace element array CGH of the FLNA locus suggests distinct molecular mechanisms for each of these rearrangements and implicates nearby low-copy repeats in their pathogenesis.
Conclusions:
These results demonstrate that FLNA is prone to pathogenic rearrangements and highlight the importance of screening for CNV in people with PNH who lack FLNA point mutations. Neurology® 2012;78:269–278
Congenital disorders of human brain development represent a clinically and genetically diverse group of conditions. Loss-of-function mutations in FLNA, which encode the actin cross-link protein Filamin A, cause one of the most prevalent brain malformations encountered clinically: X-linked periventricular nodular heterotopia (PNH).1–3 In PNH, the neurons accumulate as nodules along the surface of the lateral ventricles,1 leading to seizures in early adulthood and sometimes preventing normal psychomotor development.1 Because loss-of-function mutations in FLNA are usually prenatally lethal in males, X-linked PNH is found clinically primarily in females.1,4
FLNA point mutations are found in most, but not all, patients with familial X-linked PNH,5,6 and in only 26% of sporadic patients with bilateral PNH.5 Some of the missing heritability may This may be due to genetic heterogeneity.7–10 Alternatively, Filamin A lesions undetectable by traditional sequencing can also cause disease. One category of such lesions is copy number variants (CNVs), which are an increasingly recognized cause of disease11,12 and trait variation.13–15 FLNA resides in a region prone to X-chromosome rearrangement at Xq28.16,17 that CNVs in this region could represent a previously unrecognized cause of PNH.
Here, we describe 3 patients with pathogenic FLNA CNV as a cause of unexplained PNH. Custom mosaic oligo comparative genomic hybridization (CGH) revealed distinct CNV boundaries consistent with rare and non-recurrent events, likely stimulated by nearby genomic low copy repeats (LCRs). These results demonstrate that genomic rearrangements are an important cause of FLNA mutation and establish the clinical importance of screening for such rearrangements in PNH.